Topology evaluation of models for difficult targets in the 14th round of the critical assessment of protein structure prediction - Kinch - - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library
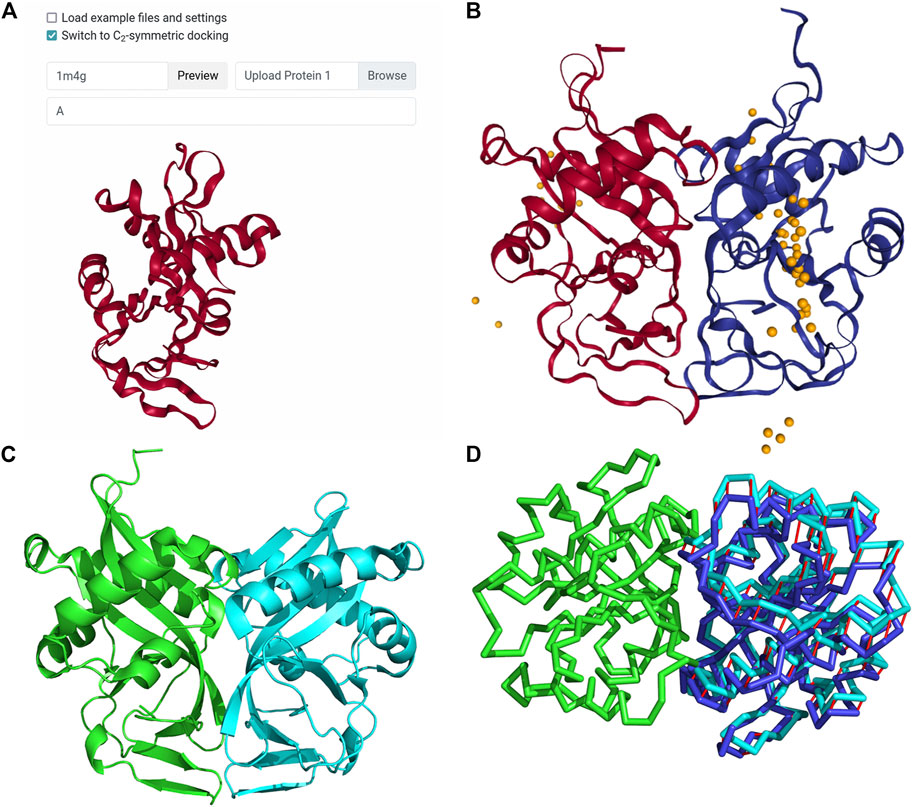
Frontiers | LZerD Protein-Protein Docking Webserver Enhanced With de novo Structure Prediction | Molecular Biosciences

Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation‐induced damage | The EMBO Journal

Examples of successful QUARK modeling results on known Hard E. coli... | Download Scientific Diagram
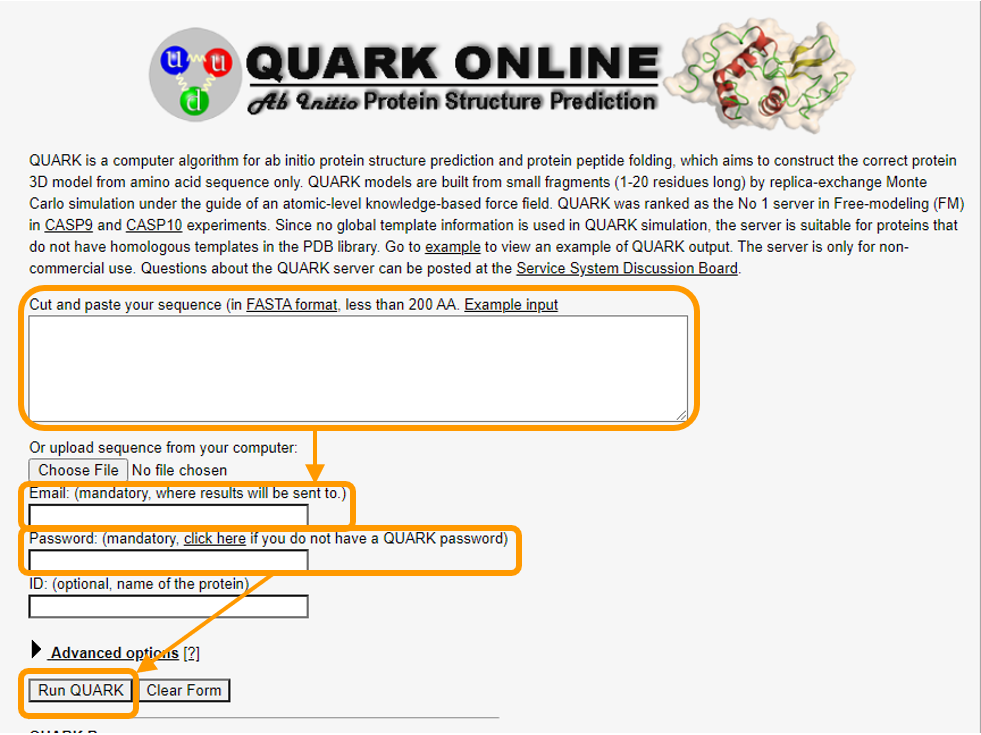
Software Tutorial: Using ab initio Modeling to Predict the Structure of Hemoglobin Subunit Alpha - Biological Modeling

Overview of the RBO Aleph structure prediction pipeline: the server... | Download Scientific Diagram
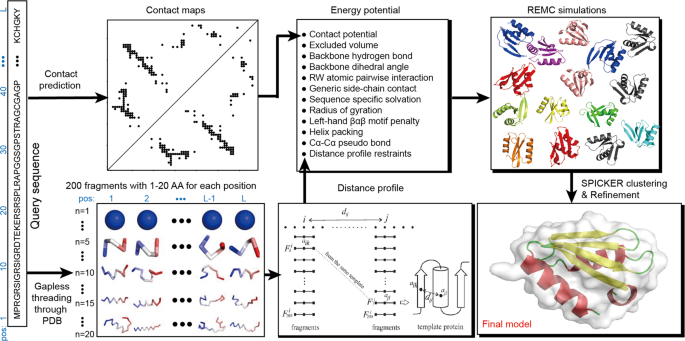
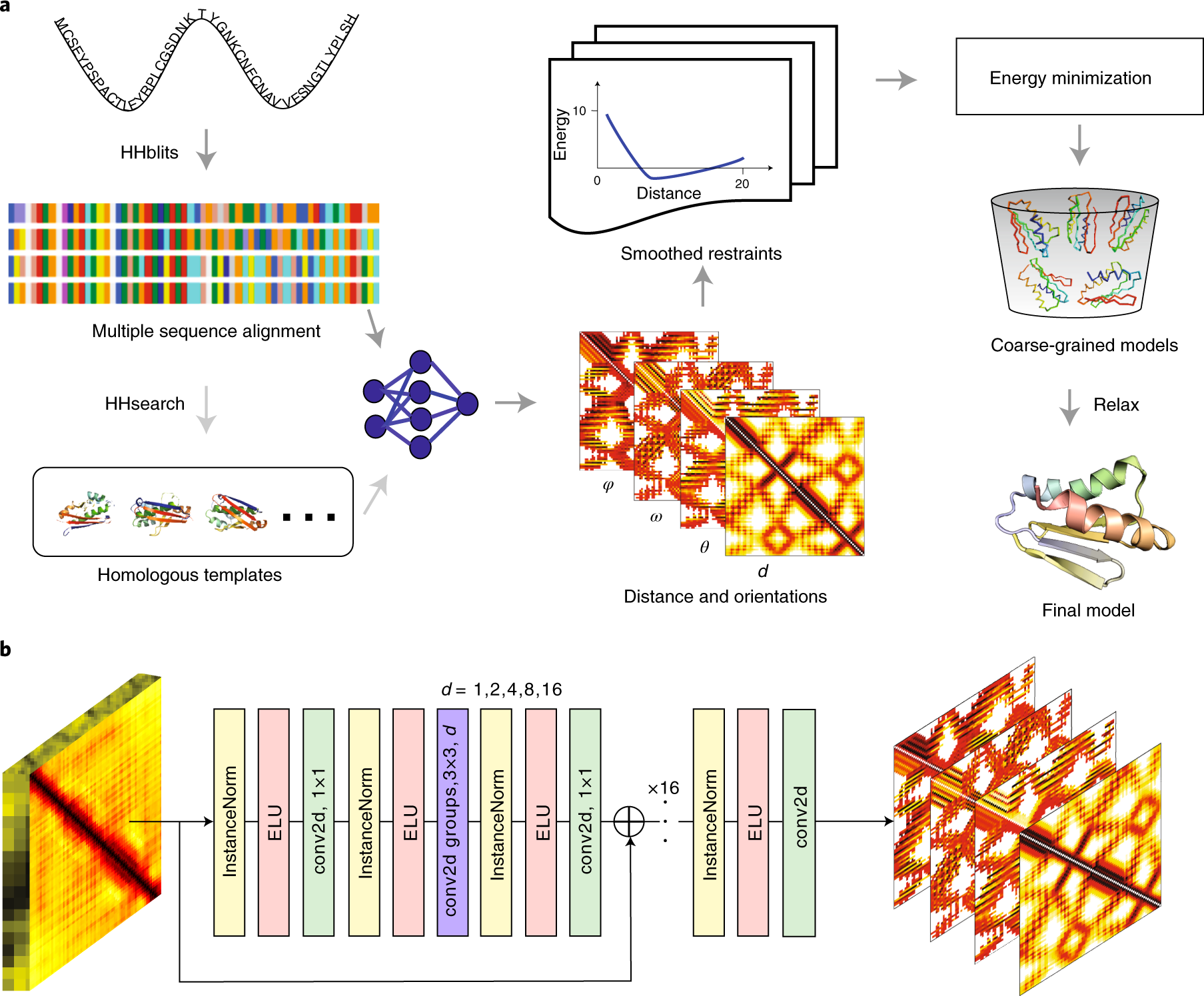
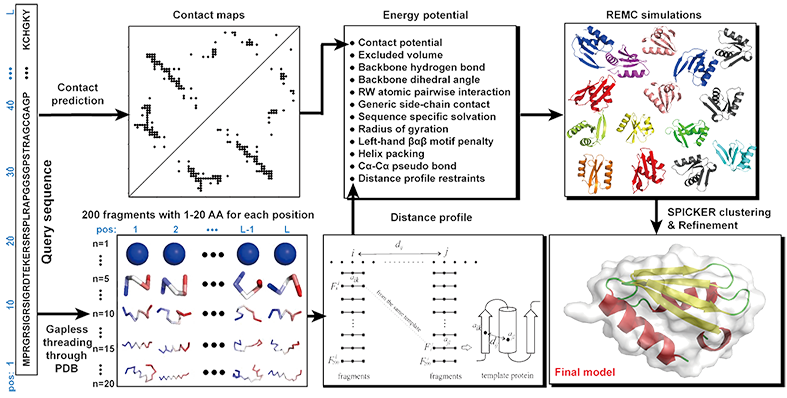
Improving fragment-based ab initio protein structure assembly using low-accuracy contact-map predictions | Nature Communications
![PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/682fc6a2dcf6edaadacb1a2f90fdd137aef28d15/2-Figure1-1.png)
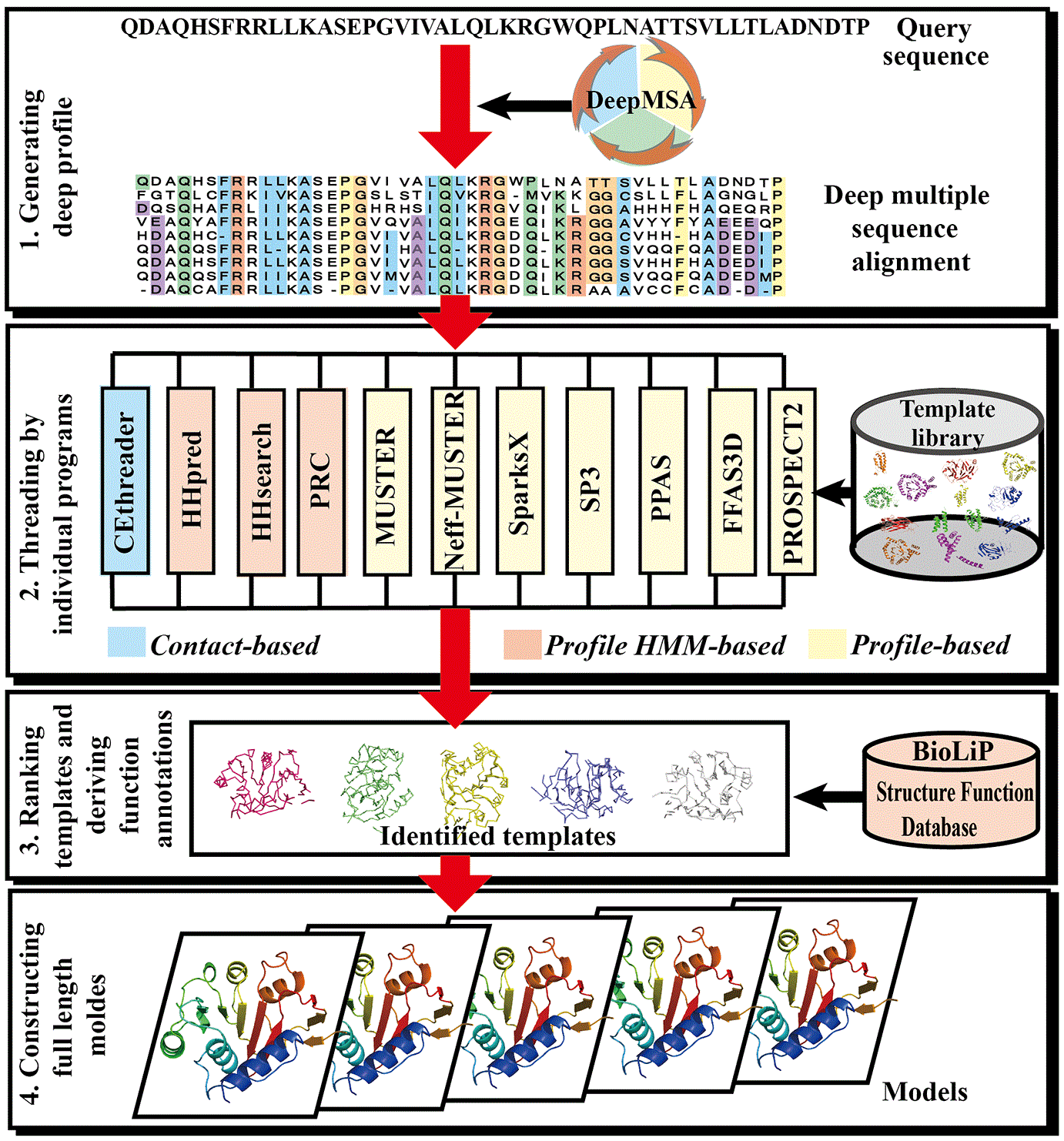
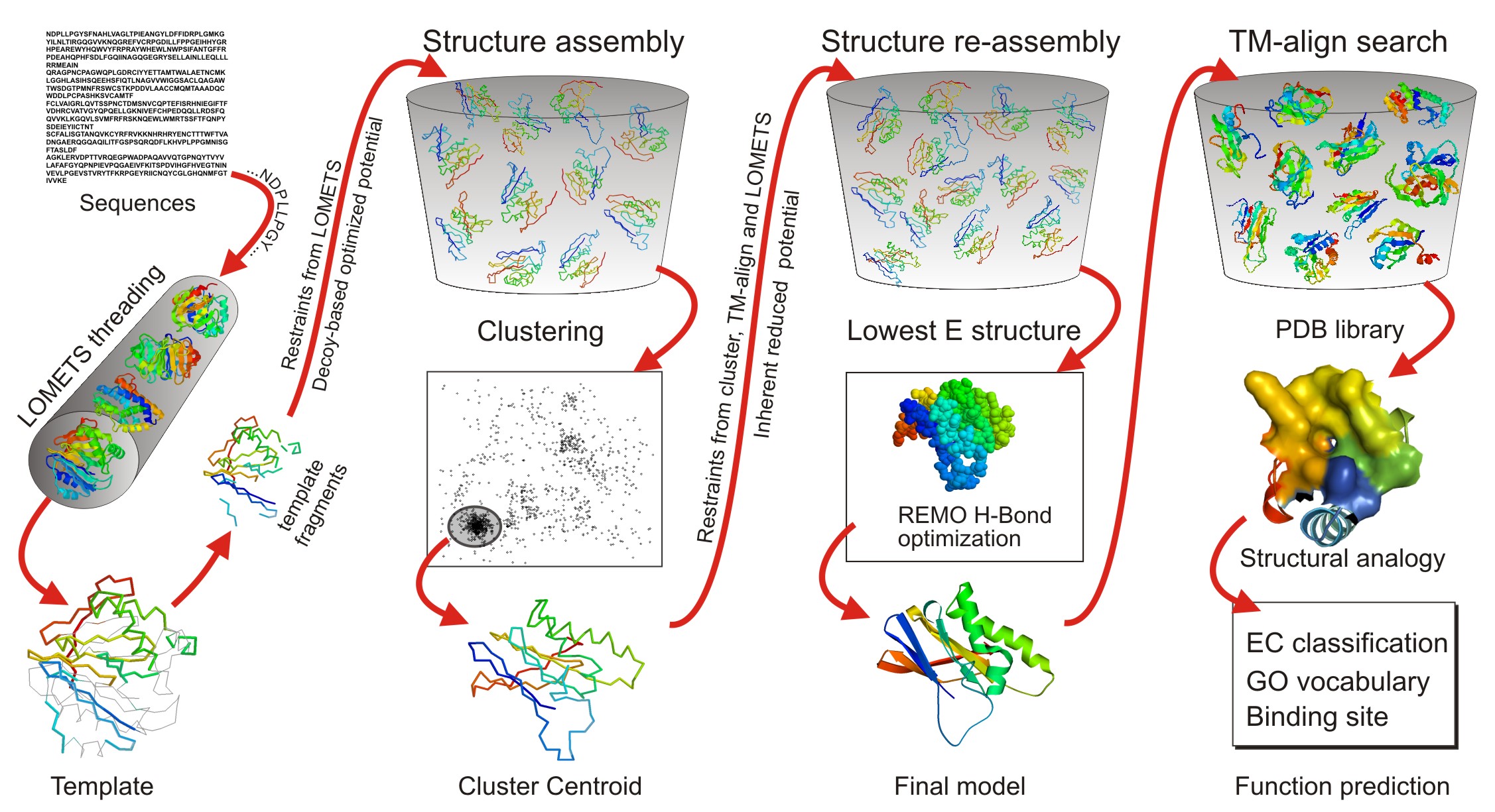
PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar

I-TASSER gateway: A protein structure and function prediction server powered by XSEDE - ScienceDirect

Secondary structure alignment of CsgA models. In (a), the percentage of... | Download Scientific Diagram
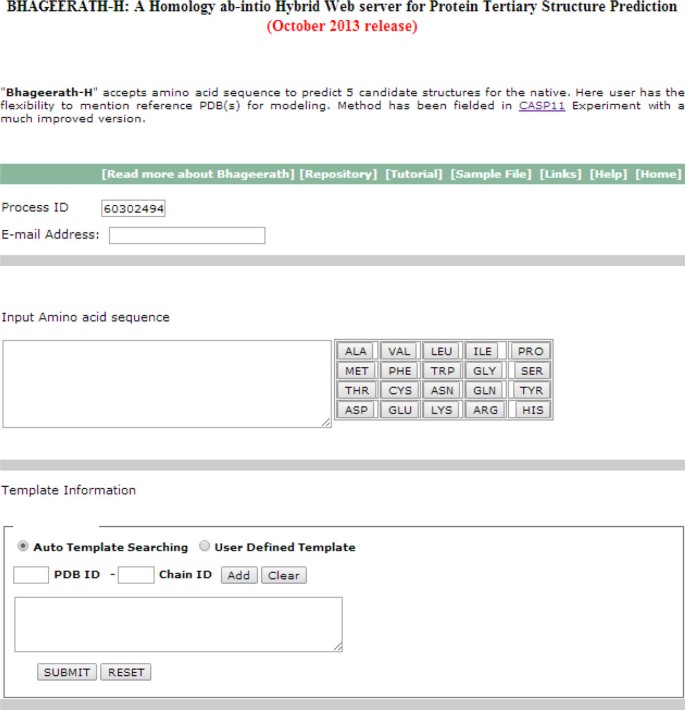
Bhageerath-H: A homology/ab initio hybrid server for predicting tertiary structures of monomeric soluble proteins | BMC Bioinformatics | Full Text
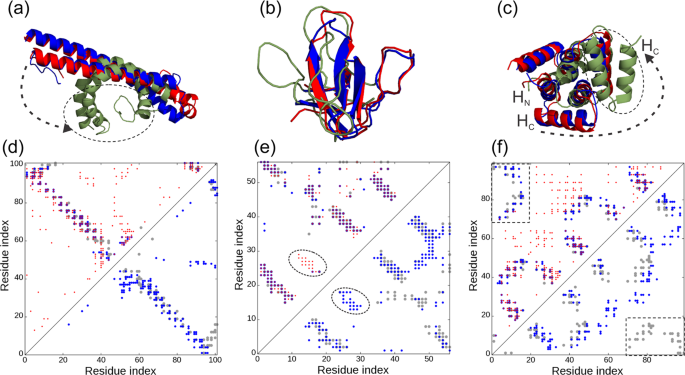
Improving fragment-based ab initio protein structure assembly using low-accuracy contact-map predictions | Nature Communications

Interplay of I‐TASSER and QUARK for template‐based and ab initio protein structure prediction in CASP10 - Zhang - 2014 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library